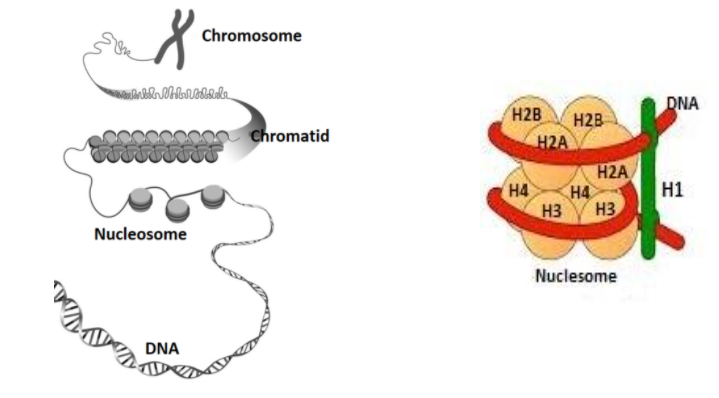
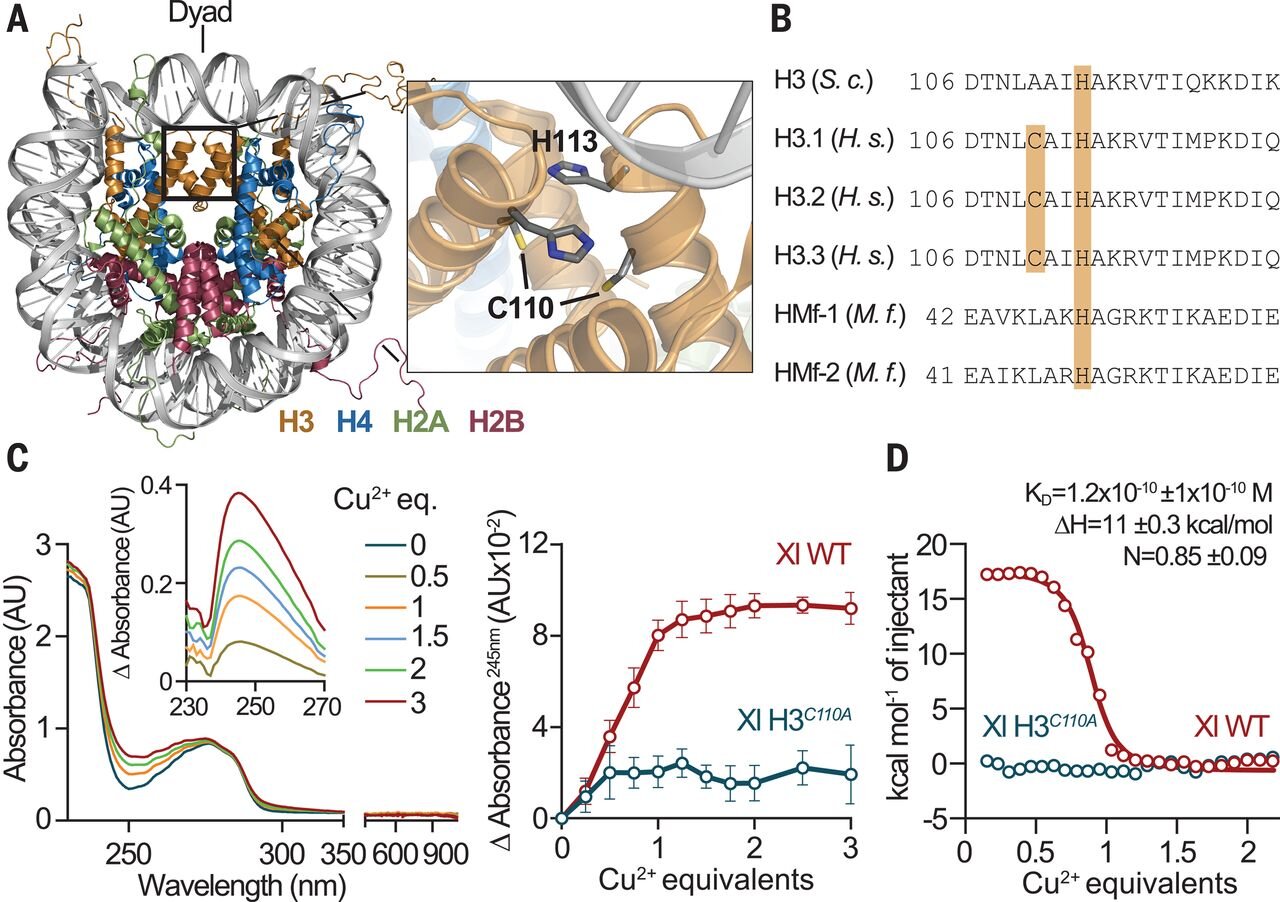
POLE3-POLE4 Is a Histone H3-H4 Chaperone that Maintains Chromatin Integrity during DNA Replication - ScienceDirect

Histones H3 and H4 require their relevant amino-tails for efficient nuclear import and replication-coupled chromatin assembly in vivo | Scientific Reports
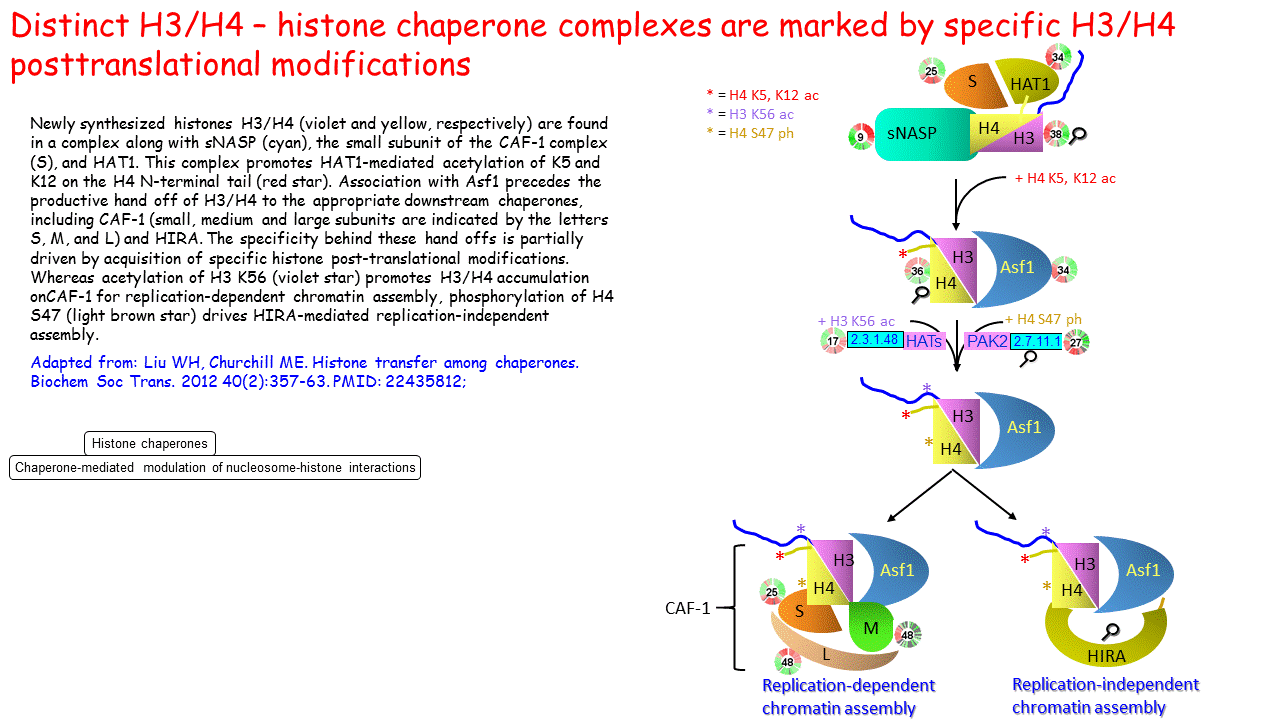
Distinct H3/H4 – histone chaperone complexes are marked by specific H3/H4 posttranslational modifications
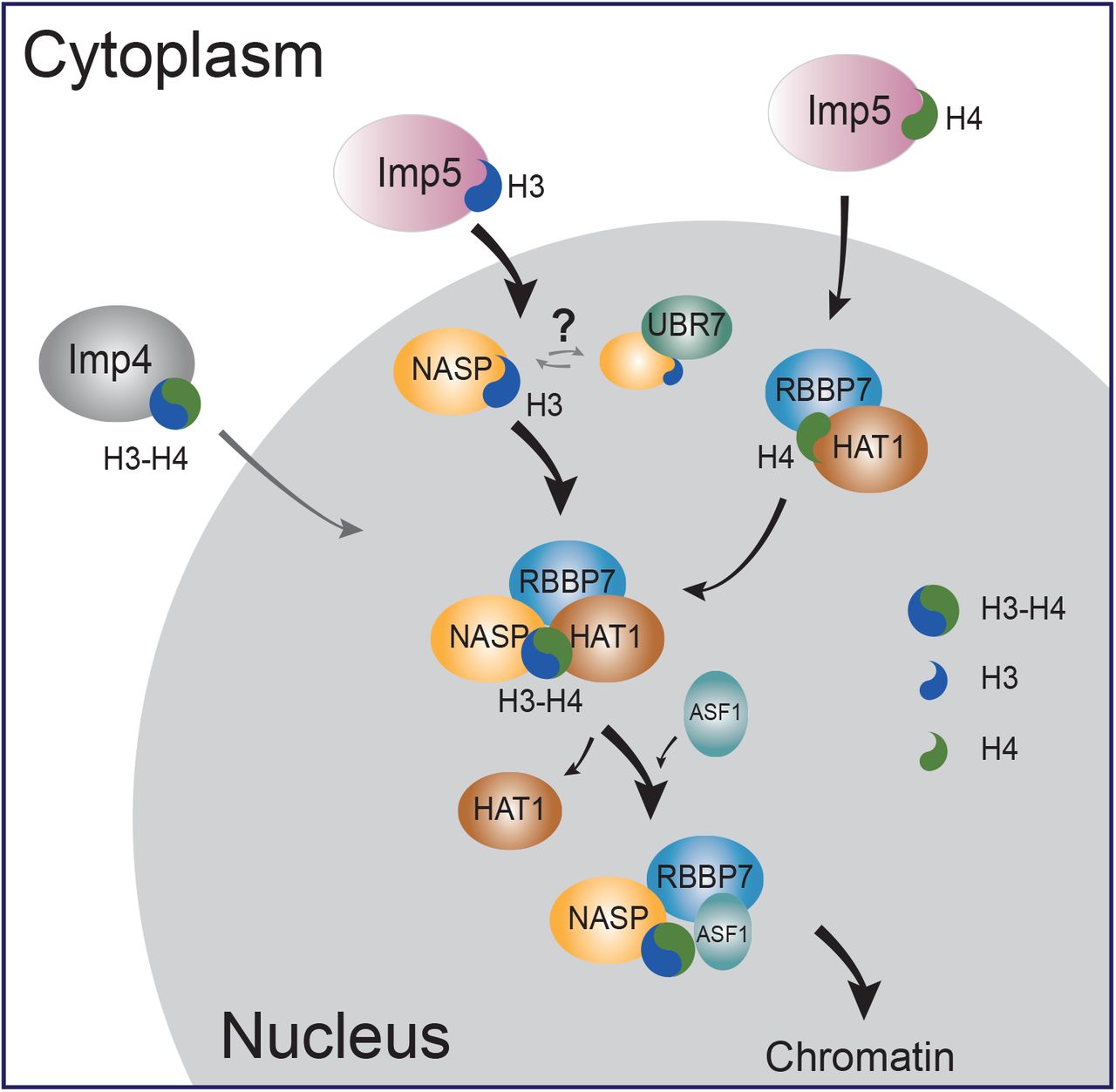
Nuclear import and chaperoning of monomeric histones H3 and H4 is mediated by Imp5, NASP and the HAT1 complex | bioRxiv

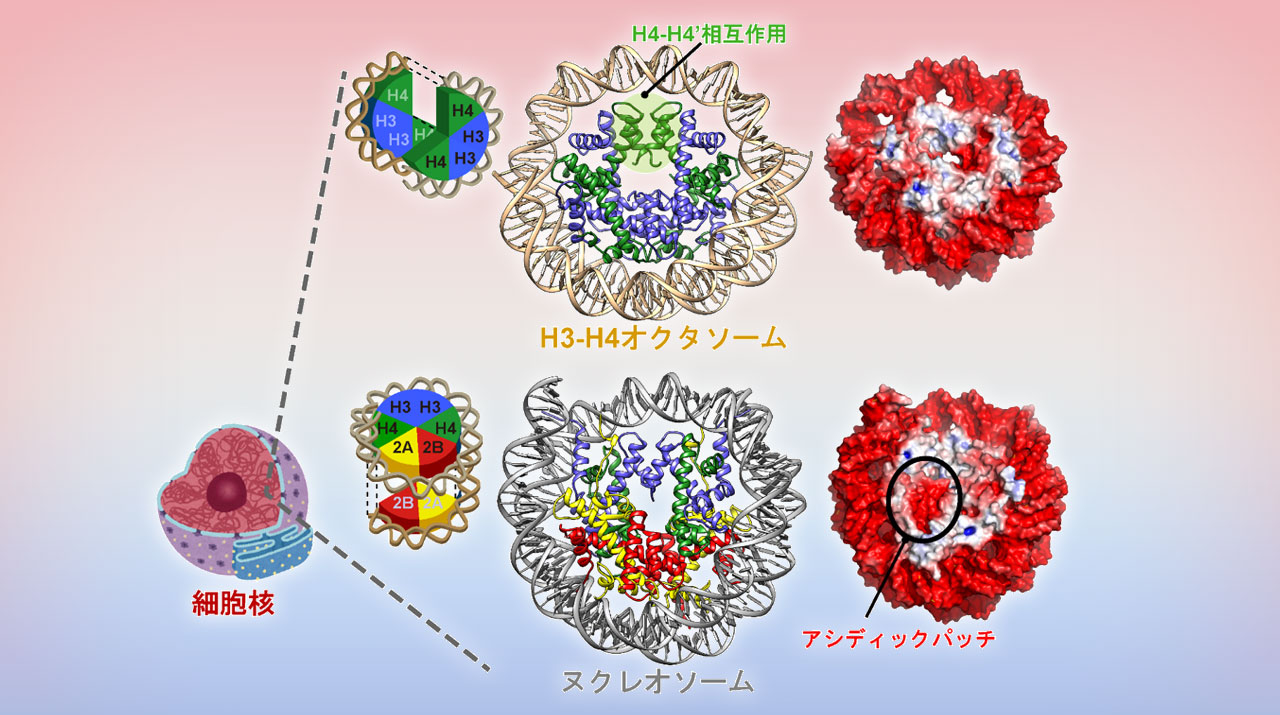
Cryo–electron microscopy structure of the H3-H4 octasome: A nucleosome-like particle without histones H2A and H2B | PNAS

Simultaneous Proteoform Analysis of Histones H3 and H4 with a Simplified Middle-Down Proteomics Method | Analytical Chemistry

Insights into the molecular architecture and histone H3-H4 deposition mechanism of yeast Chromatin assembly factor 1 | eLife

A unique binding mode enables MCM2 to chaperone histones H3–H4 at replication forks | Nature Structural & Molecular Biology

Distinct H3/H4 – histone chaperone complexes are marked by specific... | Download Scientific Diagram

The Histone Chaperones Nap1 and Vps75 Bind Histones H3 and H4 in a Tetrameric Conformation - ScienceDirect
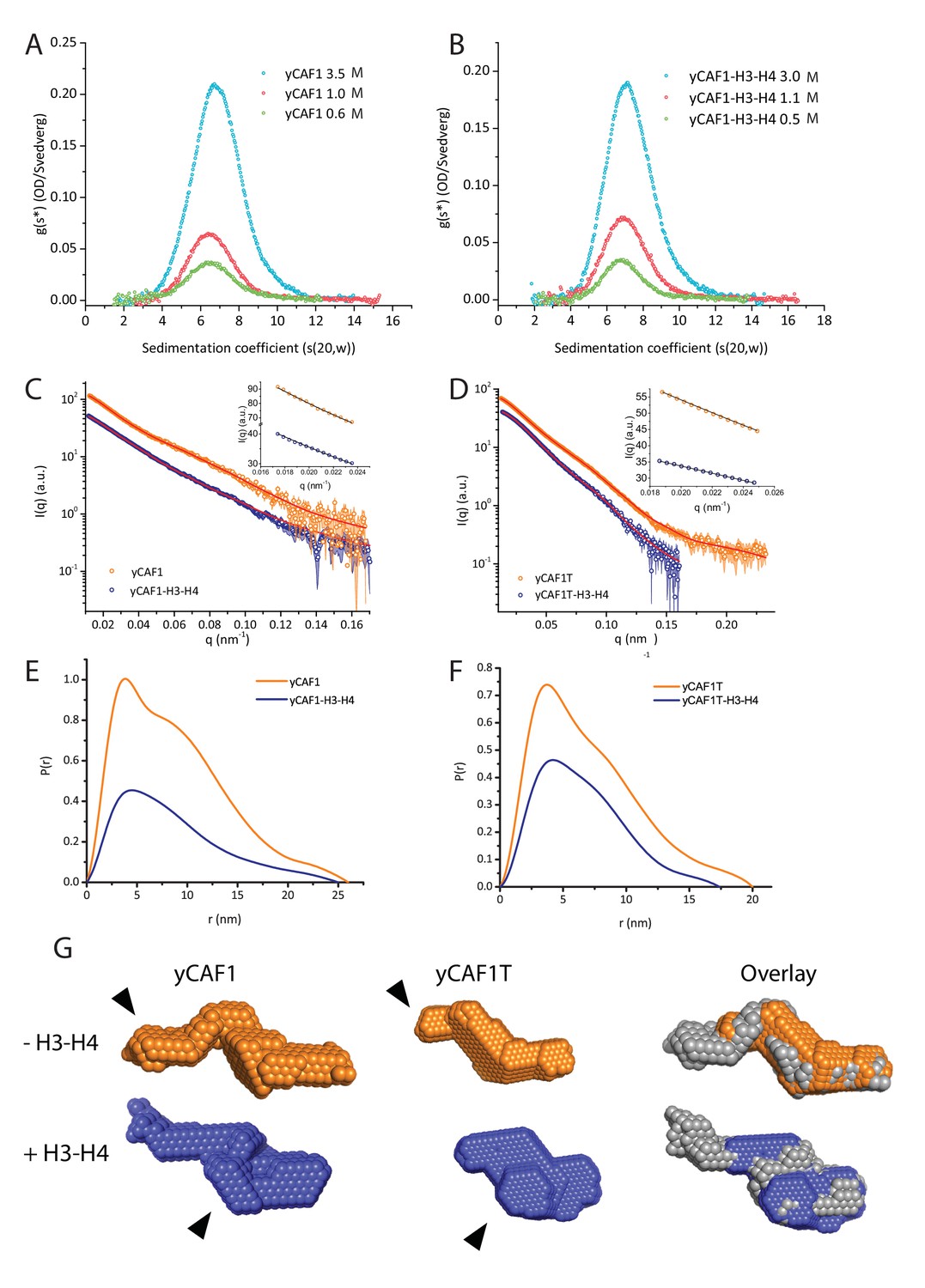
Insights into the molecular architecture and histone H3-H4 deposition mechanism of yeast Chromatin assembly factor 1 | eLife